Dimensionality Reduction: PCA and NMF
High-dimensional datasets
Dealing with many variables
- So far we’ve largely concentrated on cases in which we have relatively large numbers of measurements for a few variables
- This is frequently referred to as \(n > p\)
- Two other extremes are important
- Many observations and many variables
- Many variables but few observations (\(p > n\))
Why many variables are hard
Usually, when we’re dealing with many variables, we don’t have a great understanding of how they relate to each other.
- E.g. if gene X is high, we can’t be sure that will mean gene Y will be too
- If we had these relationships, we could reduce the data
- E.g. if we had variables to tell us it’s 3 pm in Los Angeles, we don’t need one to say it’s daytime
Dimensionality Reduction
Generate a low-dimensional encoding of a high-dimensional space
Purposes:
- Data compression / visualization
- Robustness to noise and uncertainty
- Potentially easier to interpret
Bonus: Many of the other methods from the class can be applied after dimensionality reduction with little or no adjustment!
Matrix factorization
Many dimensionality reduction methods involve matrix factorization
Basic Idea: Find two (or more) matrices whose product best approximates the original matrix
Low rank approximation to original \(N\times M\) matrix:
\[ \mathbf{X} \approx \mathbf{W} \mathbf{H}^\top \]
where \(\mathbf{W}\) is \(N\times R\), \(\mathbf{H}\) is \(M\times R\), and \(R \ll \min(N, M)\).
Visualizing matrix factorization
Visualization of matrix factorization, showing a data matrix X approximated by the product of W and H transpose, with sample and feature axes and common names for the factor matrices in PCA and NMF.
Generalization of many methods (e.g., SVD, QR, CUR, Truncated SVD, etc.)
How to choose the rank, \(R\)?
\[ \mathbf{X} \approx \mathbf{W} \mathbf{H}^\top \]
where \(\mathbf{W}\) is \(N\times R\), \(\mathbf{H}\) is \(M\times R\), and \(R \ll \min(N, M)\).
Example: image compression
Original image

https://www.aaronschlegel.com/image-compression-principal-component-analysis/
Rank = 3

https://www.aaronschlegel.com/image-compression-principal-component-analysis/
Rank = 46

https://www.aaronschlegel.com/image-compression-principal-component-analysis/
Applications
Process control
- Large bioreactor runs may be recorded in a database, along with a variety of measurements from those runs
- We may be interested in how those different runs varied, and how each factor relates to one another
- Plotting a compressed version of that data can indicate when an anomalous change is present
Mutational processes
- Anytime multiple contributory factors give rise to a phenomenon, matrix factorization can separate them out
- Will talk about this in greater detail
Cell heterogeneity
- Enormous interest in understanding how cells are similar or different
- Answer to this can be in millions of different ways
- But cells often follow programs
Principal Components Analysis (PCA)
PCA as matrix factorization
For data table \(\mathbf{X}\) with its singular value decomposition, \(\mathbf{X} = \mathbf{U}\mathbf{D}\mathbf{V}^\top\),
- \(\mathbf{W} = \mathbf{U}\mathbf{D}\) gives the scores, placing observations in a low-dimensional space
- \(\mathbf{H} = \mathbf{V}\) gives the loadings, relating the features to that low-dimensional space
- Each principal component (PC) is a linear combination of the original features
- PCs are uncorrelated with each other
PCA intuition
- Ordered in terms of variance
- \(k\)th PC is orthogonal to all previous PCs
- Reduce dimensionality while maintaining maximal variance
- PCA minimizes reconstruction error: \(\left\|\mathbf{X} - \mathbf{W}\mathbf{H}^\top\right\|_F^2\)
Before running PCA: preprocess with care
- PCA is sensitive to the scale of the variables
- Usually center each feature to mean 0 first
- Often scale each feature to unit variance when variables are in different units
- Otherwise a high-variance measurement can dominate PC1
- Any new samples must be transformed using the same centering and scaling learned from the training data
PCA example: words
- Consider an example dataset of two variables about words
- The number of lines of its dictionary definition
- The length of a word
- Example from Abdi & Williams, 2010
- We obtain \(\mathbf{X}\) after applying appropriate centering and normalization
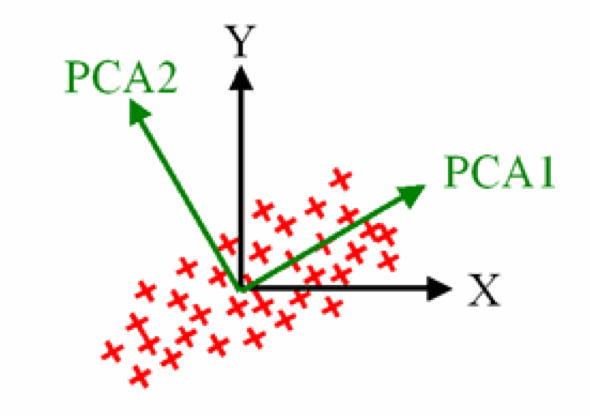
PCA identifies two orthogonal axes of variation
- The first component (PC1) explains the most variation
- The second component (PC2) is orthogonal to PC1
One can project the observed data points onto the principal components
Scores for each data point
- We can read the projected scores for each data point
Projecting new data with PCA
- Once you have characterized the PCA components, you may use the loadings to project new data points.
PCA words example: projecting a new word, ‘sur’.
Score for the new word, ‘sur’.
PCA gives a low-rank approximation
- PCA approximates the original data using only a few principal directions
- Keeping the first \(R\) PCs gives the best rank-\(R\) linear approximation in least squares terms
- Geometrically: project the data onto a lower-dimensional subspace, then reconstruct back into the original space
Visualization of PCA as a low-rank approximation: data points are approximated by projection onto the leading principal component directions.
Interpreting PCA outputs
- Scores tell you where each observation lies in PC space
- Loadings tell you which original features define each PC
- Large-magnitude loadings indicate features that contribute strongly to that component
- The sign of a PC is arbitrary
- Flipping all scores and loadings for a component gives the same model
How many PCs should we keep?
- Look at the explained variance ratio
- Use a scree plot to look for an elbow
- Keep enough PCs for the downstream goal
- Visualization may need only 2-3 PCs
- Denoising or compression may need more
- Validate the choice with reconstruction error or downstream performance when possible
Algorithms: computing PCA
- Classical PCA is often computed with an SVD or eigendecomposition
- These are deterministic for a fixed input matrix
- Iterative methods are also used in practice
- Helpful for very large matrices or when only a few PCs are needed
- NIPALS (nonlinear iterative partial least squares)
- Able to efficiently calculate a few PCs without a full decomposition
- Useful when the full SVD would be expensive
PCA in practice
- Implemented within
sklearn.decomposition.PCAPCA.fit_transform(X)fits the model toX, then provides the data in principal component spacePCA.components_provides the “loadings matrix”, or directions of maximum variancePCA.explained_variance_provides the amount of variance explained by each componentPCA.explained_variance_ratio_gives the fraction of total variance explained by each PC
PCA code example
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.decomposition import PCA
iris = datasets.load_iris()
X = iris.data
y = iris.target
target_names = iris.target_names
pca = PCA(n_components=2)
X_r = pca.fit_transform(X)
# Print PC1 loadings
print(pca.components_[0, :])
# Print PC1 scores
print(X_r[:, 0])
# Percentage of variance explained for each component
print(pca.explained_variance_ratio_)
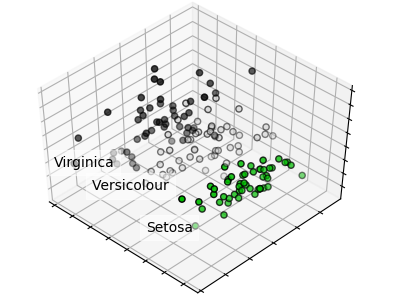
# [ 0.92461621 0.05301557]PCA example: separating flower species
PCA example: genes mirror geography within Europe
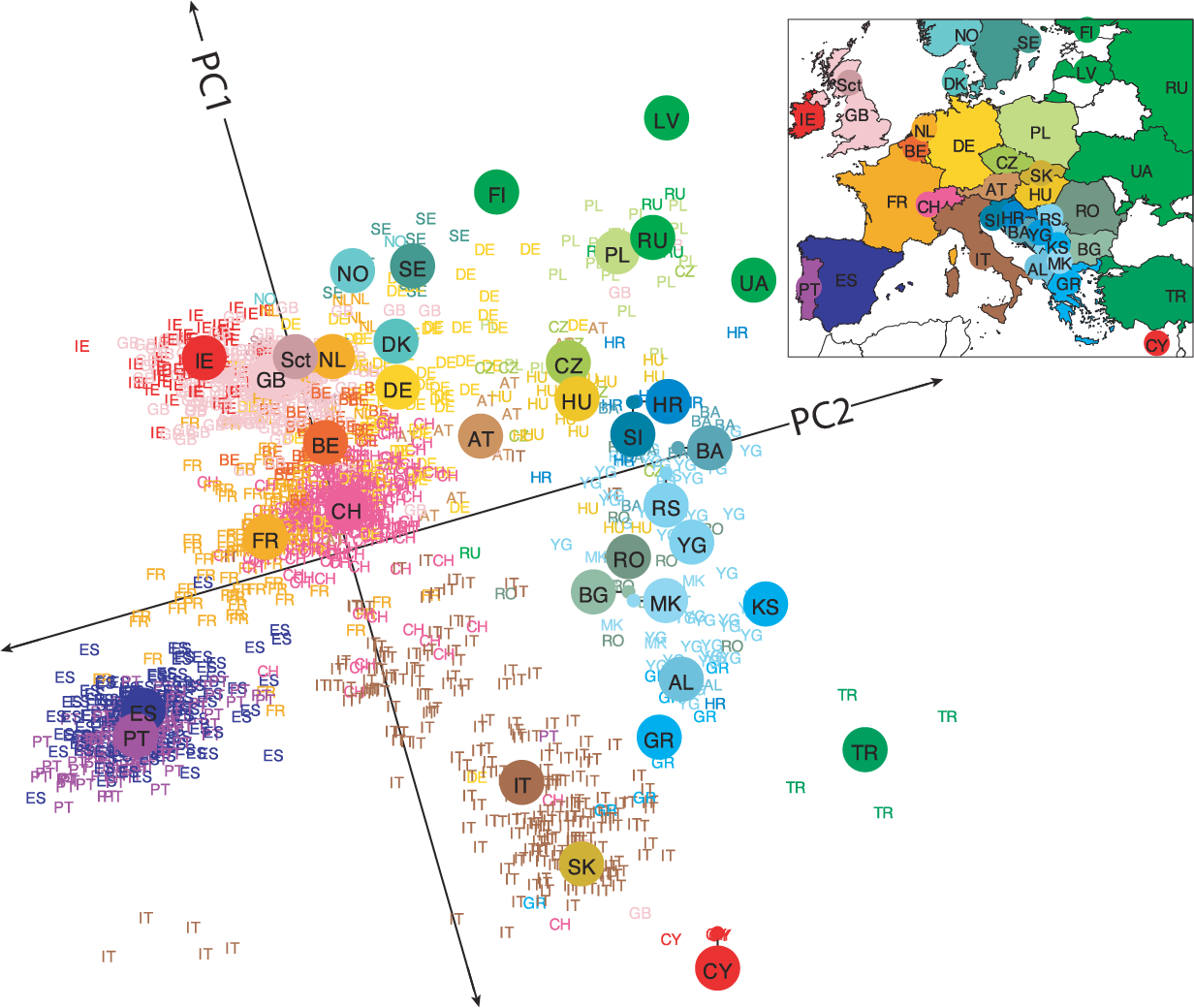
A statistical summary of genetic data from 1,387 Europeans based on principal component axis one (PC1) and axis two (PC2). Novembre et al. Nature. 2008.
Non-Negative Matrix Factorization (NMF)
What if we have data wherein effects always accumulate?
NMF application: mutational processes in cancer
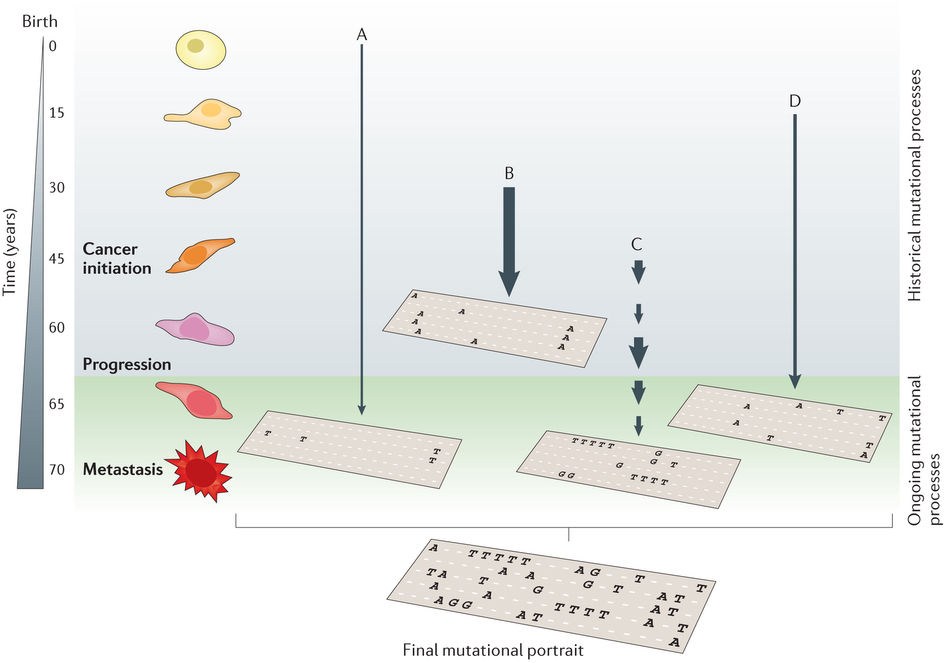
Helleday et al, Nat Rev Gen, 2014
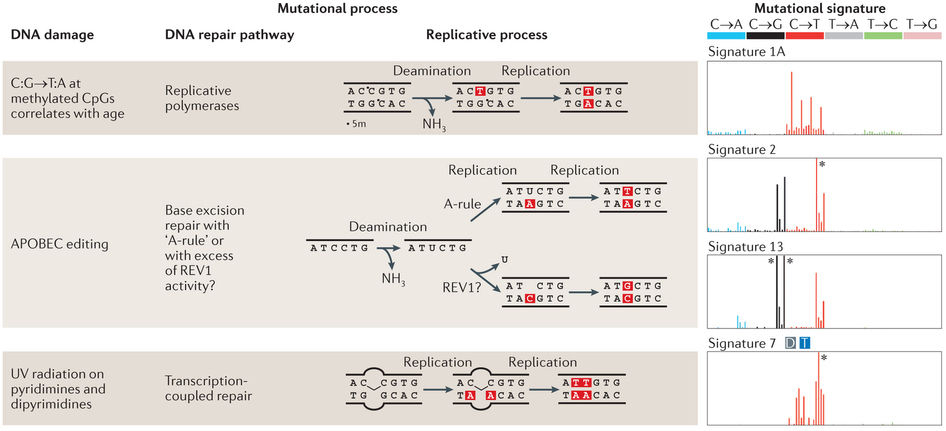
Helleday et al, Nat Rev Gen, 2014
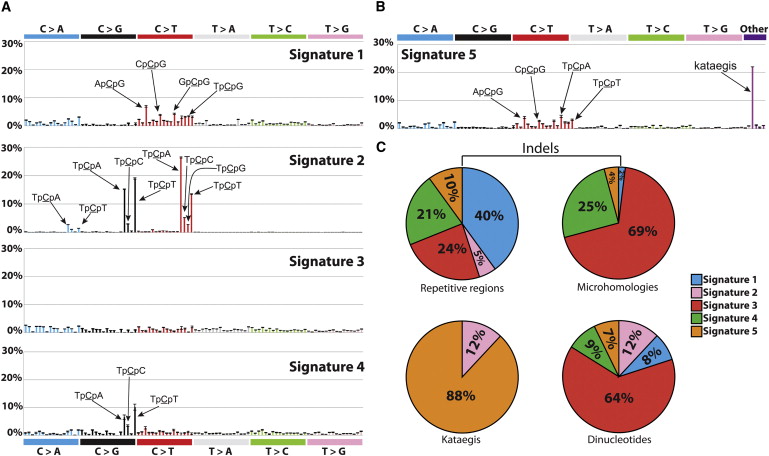
Alexandrov et al, Cell Rep, 2013
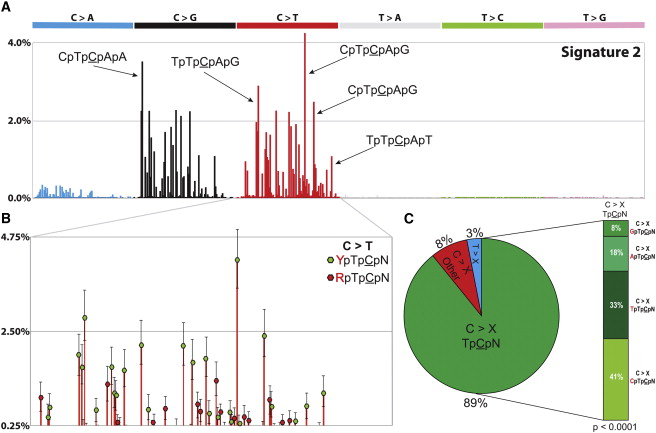
Alexandrov et al, Cell Rep, 2013
Important considerations
- Like PCA, except the coefficients must be non-negative
- Forcing positive coefficients implies an additive combination of parts to reconstruct the whole
- Leads to sparse factors
- The answer you get will always depend on the error metric, starting point, and search method
NMF objective
Find non-negative matrices \(\mathbf{W}\) and \(\mathbf{H}\) such that
\[ \mathbf{X} \approx \mathbf{W}\mathbf{H}^\top \qquad\text{with}\qquad \mathbf{W}_{ij} \ge 0,\ \mathbf{H}_{ij} \ge 0 \]
Typically, we minimize reconstruction error:
\[ \min_{\mathbf{W},\mathbf{H}} \left\|\mathbf{X} - \mathbf{W}\mathbf{H}^\top\right\|_F^2 \qquad \text{subject to } \mathbf{W}, \mathbf{H} \ge 0 \]
Multiplicative updates
- The update rule is multiplicative instead of additive
- If the initial values for \(\mathbf{W}\) and \(\mathbf{H}\) are non-negative, then \(\mathbf{W}\) and \(\mathbf{H}\) can never become negative
- This guarantees a non-negative factorization
- Will converge to a local minimum
- Therefore starting point matters
Updating W
\[[W]_{ij} \leftarrow [W]_{ij} \frac{[\color{darkred} X \color{darkblue}H \color{black}]_{ij}}{[\color{darkred}W \color{darkblue}{H^T H} \color{black}]_{ij}}\]
Color indicates the data term and the factor term.
Updating H
\[[H]_{ij} \leftarrow [H]_{ij} \frac{[\color{darkred}{X^T} \color{darkblue}W \color{black}]_{ij}}{[\color{darkred}H \color{darkblue}{W^T W} \color{black}]_{ij}}\]
Color indicates the data term and the factor term.
Coordinate descent
- Another approach is to find the gradient across all the variables in the matrix
- Not going to go through implementation
- Will also converge to a local minimum
NMF in practice
- Implemented within
sklearn.decomposition.NMF.n_components: number of componentsinit: how to initialize the searchsolver: ‘cd’ for coordinate descent, or ‘mu’ for multiplicative updatel1_ratio,alpha_H,alpha_W: Can regularize fit
- Provides:
NMF.components_: components x features matrix- Returns transformed data through
NMF.fit_transform()
Review
Summary: PCA and NMF
PCA
- Preserves the covariation within a dataset
- Therefore mostly preserves axes of maximal variation
- Number of components will vary in practice
NMF
- Explains the dataset through two non-negative matrices
- Much more stable patterns when assumptions are appropriate
- Will explain less variance for a given number of components
- Excellent for separating out additive factors
Closing
As always, selection of the appropriate method depends upon the question being asked.
Reading & Resources
Review Questions
- What do dimensionality reduction methods reduce? What is the tradeoff?
- What are three benefits of dimensionality reduction?
- Does matrix factorization have one answer? If not, what are two choices you could make?
- What does principal components analysis aim to preserve?
- What are the minimum and maximum number of principal components one can have for a dataset of 300 observations and 10 variables?
- How can you determine the “right” number of PCs to use?
- What is a loading matrix? What would be the dimensions of this matrix for the dataset in Q5 when using three PCs?
- What is a scores matrix? What would be the dimensions of this matrix for the dataset in Q5 when using three PCs?
- By definition, what is the direction of PC1?
- See question 5 on midterm W20. How does movement of the siControl EGF point represent changes in the original data?