Double Descent and the Overparameterization Paradox
Aaron Meyer
Application
Predicting Binding at Equilibrium
- A ligand \(L\) binds a receptor \(R\) to form a complex \(RL\): \[R + L \underset{k_\text{off}}{\stackrel{k_\text{on}}{\rightleftharpoons}} RL\]
- At equilibrium, the dissociation constant is: \[K_d = \frac{k_\text{off}}{k_\text{on}} = \frac{[R][L]}{[RL]}\]
- Fractional occupancy follows the saturation isotherm: \[\theta = \frac{[L]}{K_d + [L]}\]
What can we learn from equilibrium binding data?
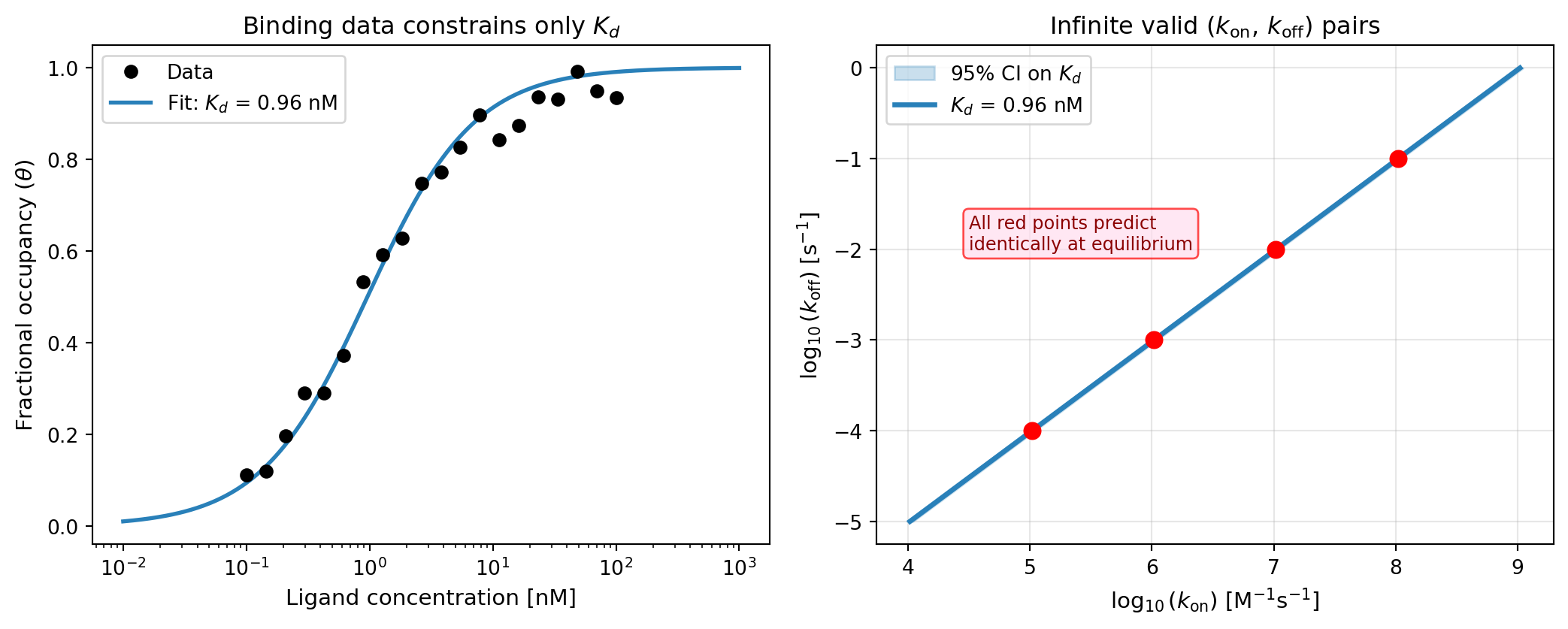
- Measuring \(\theta\) vs. \([L]\) gives us a binding curve
- Fitting the curve estimates \(K_d\), but not \(k_\text{on}\) or \(k_\text{off}\) individually
- Infinitely many \((k_\text{on}, k_\text{off})\) pairs are consistent with any measured \(K_d\): \[k_\text{off} = K_d \cdot k_\text{on} \quad \Rightarrow \quad \text{a 1-D manifold in parameter space}\]
- Yet every point on this manifold makes identical equilibrium predictions
The Classical Bias-Variance Picture
Review: Prediction Error Decomposition
\[\mathrm{PE}(\mathbf{z}_0) = \sigma_\epsilon^2 + \mathrm{Bias}^2\!\left(\hat{f}(\mathbf{z}_0)\right) + \mathrm{Var}\!\left(\hat{f}(\mathbf{z}_0)\right)\]
- Bias: systematic error—does our model class contain the true function?
- Variance: sensitivity to which training samples we happened to draw
- Classical advice: choose model complexity to trade off bias against variance
The Classical U-Curve
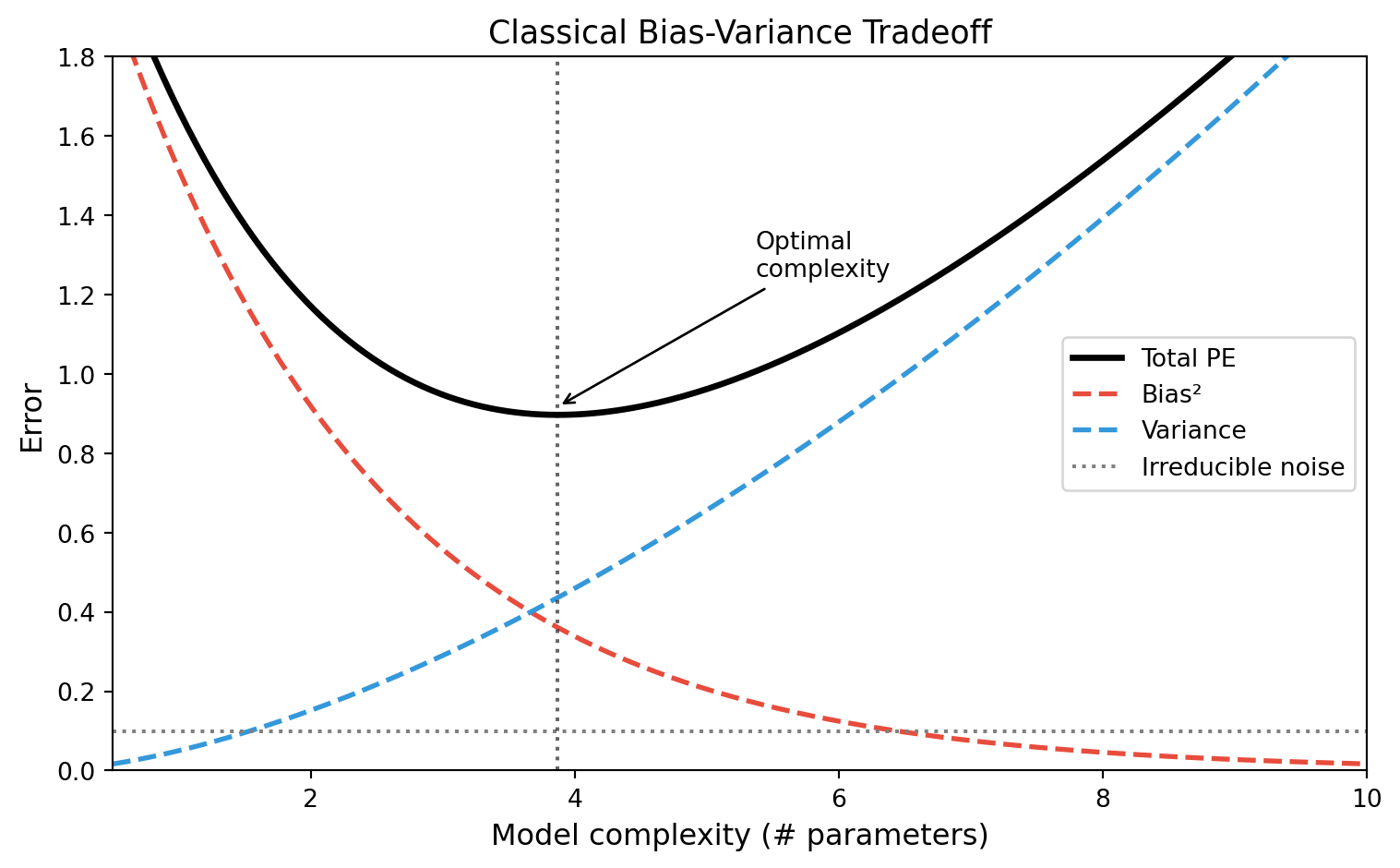
What does classical theory predict for very large models?
- As \(p \to \infty\) (more parameters than data points \(n\)), the model can fit the training data exactly
- This is called interpolation: training error \(\to 0\)
- Classical wisdom: interpolating models overfit catastrophically
- Variance \(\to \infty\) at the interpolation threshold
- Prediction error should blow up
Question: Is this always true?
The Interpolation Threshold
When \(p = n\): Exact Determination
- With exactly \(p = n\) parameters and full-rank design matrix, there is a unique solution that interpolates the data
- This solution can have very large coefficients—high variance
- Test error peaks here: the model has just enough freedom to memorize noise
When \(p > n\): Underdetermined Systems
- Infinitely many parameter vectors \(\hat{\boldsymbol{\theta}}\) satisfy \(\mathbf{X}\hat{\boldsymbol{\theta}} = \mathbf{y}\)
- Standard solvers (e.g.,
numpy.linalg.lstsq) return the minimum-norm solution: \[\hat{\boldsymbol{\theta}}^+ = \mathbf{X}^\top (\mathbf{X}\mathbf{X}^\top)^{-1} \mathbf{y}\] - Among all interpolating solutions, this is the one with smallest \(\|\boldsymbol{\theta}\|_2\)
Connection to the binding example
- Recall: from binding data, the data constrains \(K_d = k_\text{off}/k_\text{on}\), a 1-D subspace of 2-D parameter space
- The minimum-norm solution picks the shortest vector in this subspace: physically, the point where \(k_\text{on}\) and \(k_\text{off}\) are both smallest
- But all solutions make the same equilibrium predictions
| Analogy | Binding | Machine Learning |
|---|---|---|
| Parameters | \((k_\text{on}, k_\text{off})\) | Weights \(\boldsymbol{\theta}\) |
| Constraint | \(k_\text{off}/k_\text{on} = K_d\) | \(\mathbf{X}\boldsymbol{\theta} = \mathbf{y}\) |
| Prediction | \(\theta = [L]/(K_d + [L])\) | \(\hat{y} = \mathbf{x}^\top \boldsymbol{\theta}\) |
| Solution space | 1-D manifold | Affine subspace |
Double Descent
Extending the Risk Curve
- Belkin et al. (2019) demonstrated that for many model classes, the risk curve has two descents separated by a peak at the interpolation threshold
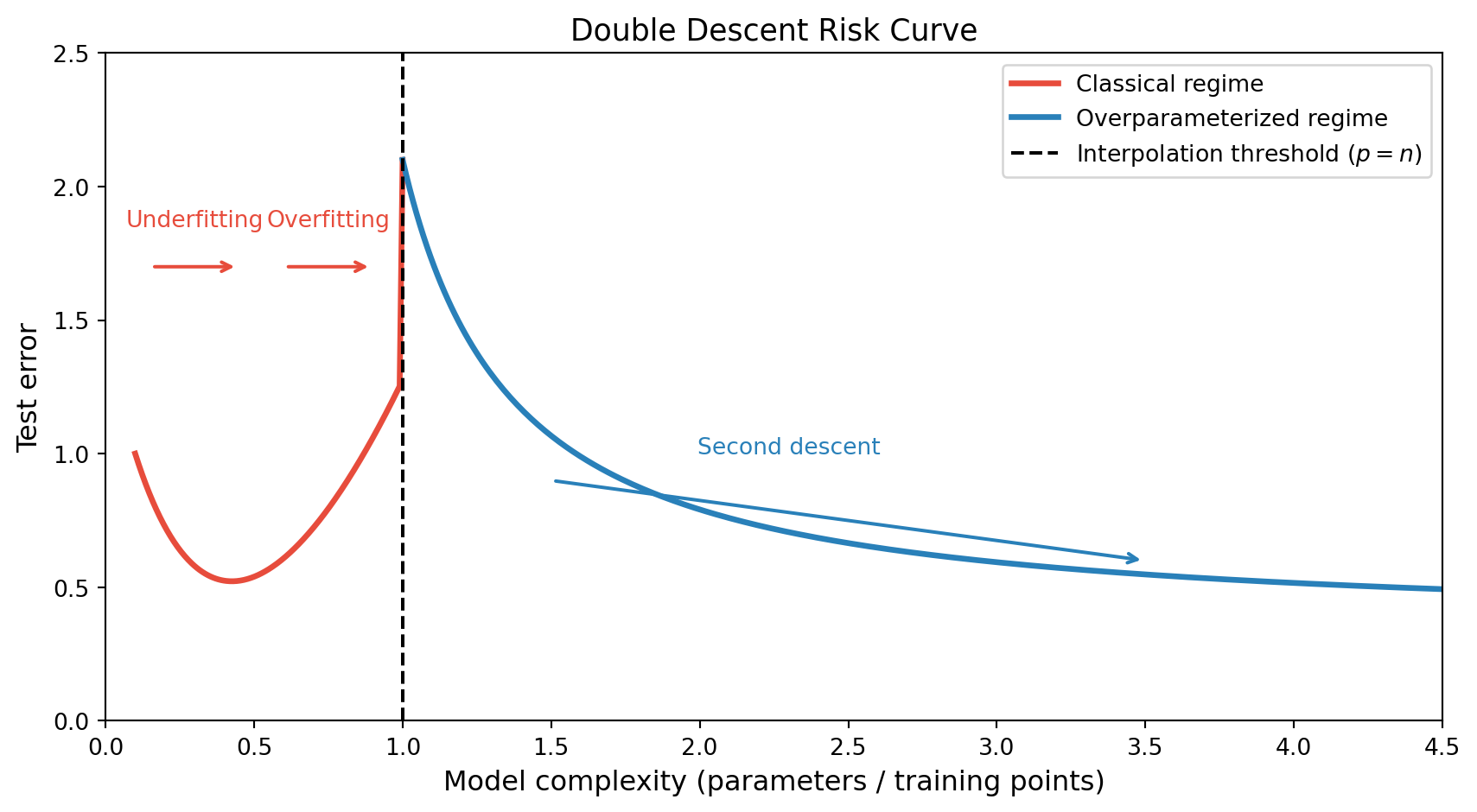
Observing double descent empirically
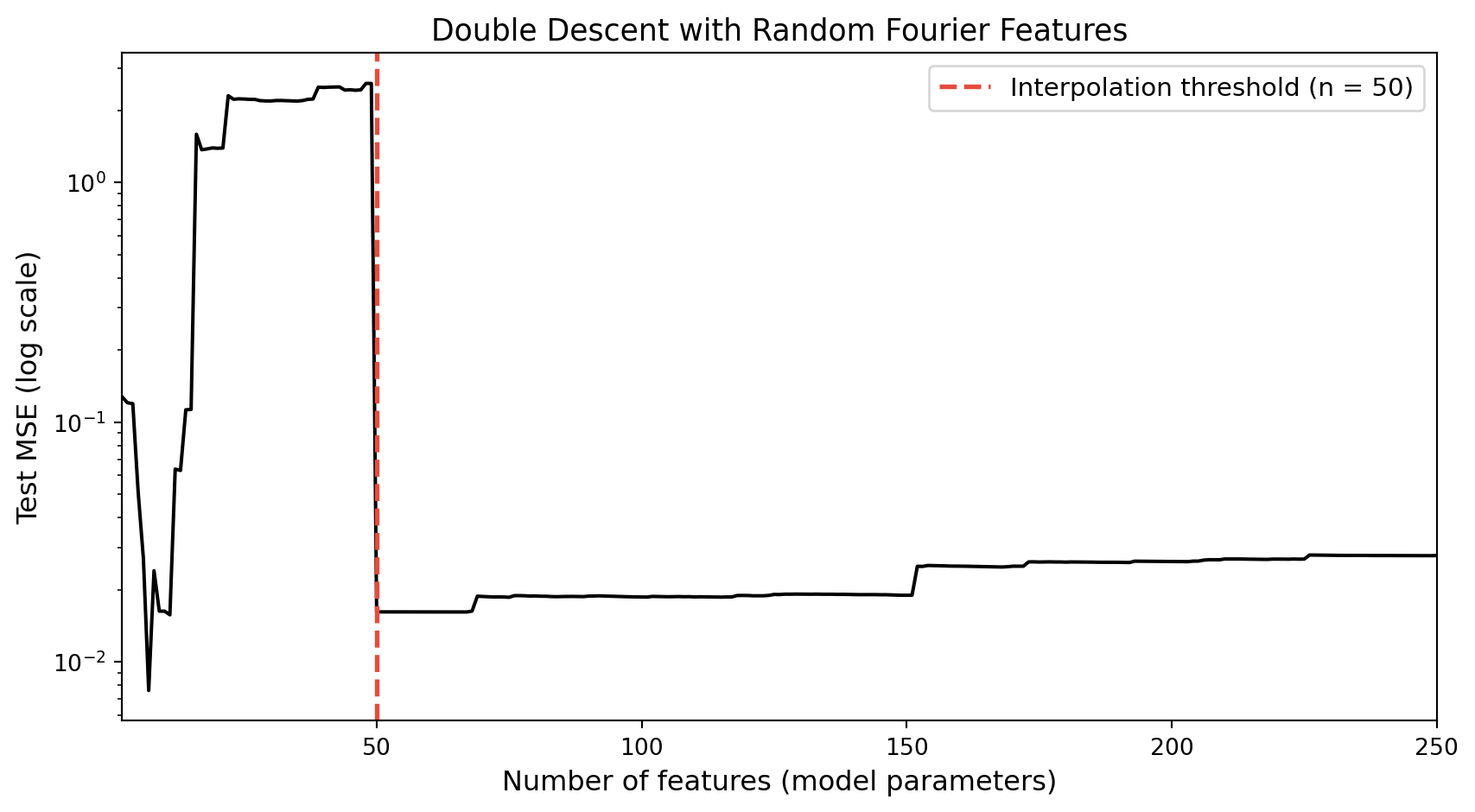
Piecewise linear fits at increasing K
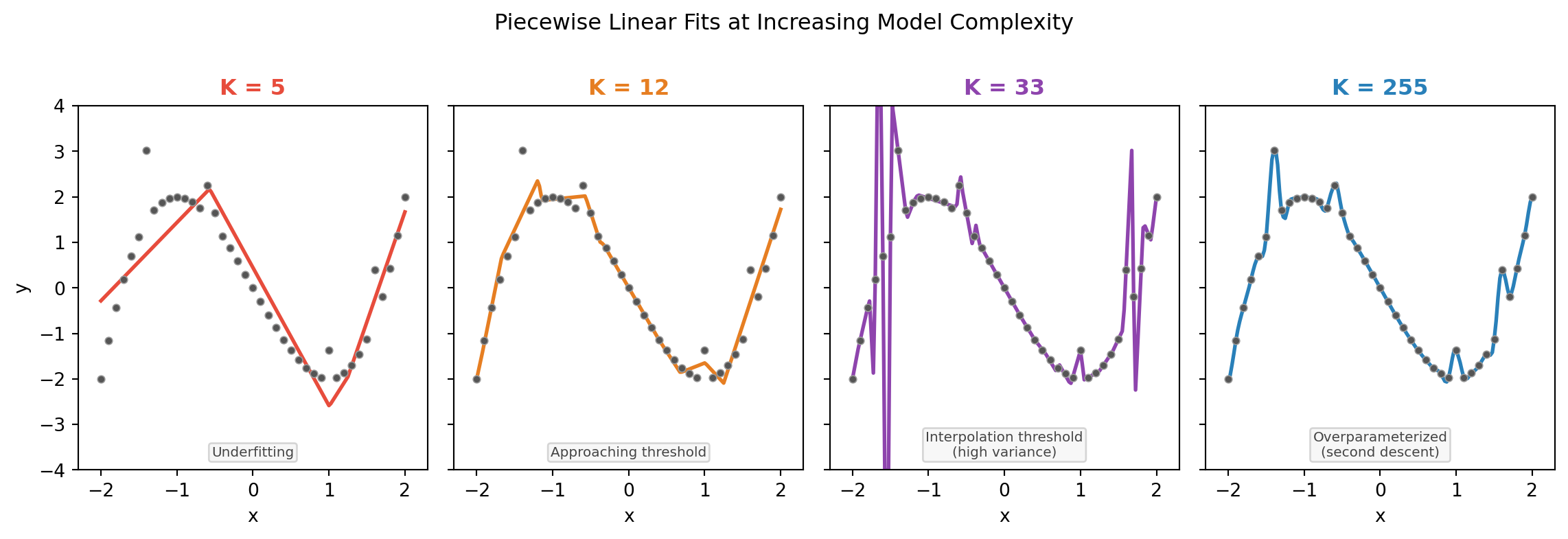
Why Does the Second Descent Happen?
- In the overparameterized regime, there are infinitely many interpolating solutions
- The minimum-norm solution (returned by gradient descent / pseudoinverse) has an implicit regularization property:
- It minimizes \(\|\boldsymbol{\theta}\|_2\) subject to fitting the training data
- This is equivalent to ridge regression in the limit \(\lambda \to 0^+\)
- It does not fit the noise dimensions beyond what is needed
\[\hat{\boldsymbol{\theta}}^+ = \arg\min_{\boldsymbol{\theta}} \|\boldsymbol{\theta}\|_2 \quad \text{s.t.} \quad \mathbf{X}\boldsymbol{\theta} = \mathbf{y}\]
Implicit regularization connects back to binding
- In the binding system, the minimum-norm solution is the \((k_\text{on}, k_\text{off})\) pair closest to the origin consistent with the measured \(K_d\)
- All solutions on the manifold predict equally—the min-norm solution does not “waste” parameter magnitude on unobserved directions
- In ML, overparameterized models similarly distribute weight across many directions; the minimum-norm solution is the smoothest interpolant
Universal Approximation
From Linear to Nonlinear Models
- Everything above applies to linear models (linear in features)
- Neural networks are nonlinear: the features themselves are learned from data
- The same overparameterization argument extends—but with far greater expressive power
The Universal Approximation Theorem
Theorem (Hornik et al., 1989; Cybenko, 1989): A feedforward network with a single hidden layer containing a sufficient number of neurons with a non-polynomial activation function can approximate any continuous function on a compact subset of \(\mathbb{R}^n\) to arbitrary precision.
- This means: given enough neurons, a neural network can represent any target function
- However, it says nothing about:
- How many neurons are “enough”
- Whether gradient descent will find the right weights
- Whether the model will generalize from finite data
A preview: overparameterized neural networks also show double descent
- Modern neural networks are massively overparameterized (often \(p \gg n\) by factors of \(10^3\)–\(10^6\))
- Like linear models, they interpolate training data and still generalize
- The implicit regularization comes from:
- Stochastic gradient descent (SGD) dynamics
- Early stopping
- Architecture choices
- Next lecture: what does it take for neural networks to actually work?
Review
Further Reading
- Belkin M. et al. Reconciling modern machine-learning practice and the bias-variance trade-off. PNAS, 2019.
- Bartlett P. et al. Benign overfitting in linear regression. PNAS, 2020.
- Zhang C. et al. Understanding deep learning requires rethinking generalization. ICLR, 2017.
- Hastie T. et al. Surprises in High-Dimensional Ridgeless Least Squares Interpolation. Ann. Statist., 2022.
Review Questions
- In the binding kinetics example, we cannot determine \(k_\text{on}\) and \(k_\text{off}\) individually from equilibrium data. Why can we still make accurate predictions about binding occupancy?
- What is the interpolation threshold? What does classical bias-variance theory predict happens to test error there?
- Describe the double descent phenomenon in words. How does it contradict the classical picture?
- What is the minimum-norm solution, and why does gradient descent find it for overparameterized linear models?
- How is the minimum-norm solution related to ridge regression? What happens as \(\lambda \to 0\)?
- A colleague says: “Your model has more parameters than training points—it must be overfitting.” How would you respond using the concepts from this lecture?
- The Universal Approximation Theorem says a sufficiently large network can approximate any continuous function. Why is this an existence result rather than a practical recipe?
- If a model achieves zero training error but low test error, what does this tell you about the model and the data?
- How does overparameterization in neural networks differ from overparameterization in linear models with random features?
- A drug discovery team trains a neural network on 500 compounds. They argue that using a 10-million-parameter model will overfit. What questions would you ask before agreeing or disagreeing?