Fitting And Regression
Project Proposals
Overview
In this project, you have two options for the general route you can take:
- Reimplement analysis from the literature.
- New, exploratory analysis of existing data.
More details in final project guidelines.
- The proposal should be less than one page and describe the following items:
- Why the topic you chose is interesting
- Demonstrate that your project fits the criteria above
- What overall approach do you plan to take for the project and why
- Demonstrate that your project can be finished within a month, including data availability
- Estimate the difficulty of your project
- We are available to discuss your ideas whenever you are ready, and you should discuss your idea with us prior to submitting your proposal.
- Recommend an early start—the earlier you finalize a proposal the sooner you can begin the project.
Application
Paternal de novo mutations
Questions:
- Where do de novo mutations arise?
- Are there factors that influence the rate of de novo mutations from one generation to another?
Application: Paternal de novo mutations
Meiosis of a cell with 2n=4 chromosomes.
Application: Paternal de novo mutations

Kong et al, Nature, 2012
- How do you infer the relationship between parental age and the rate of de novo mutations?
de novo mutations data from trios
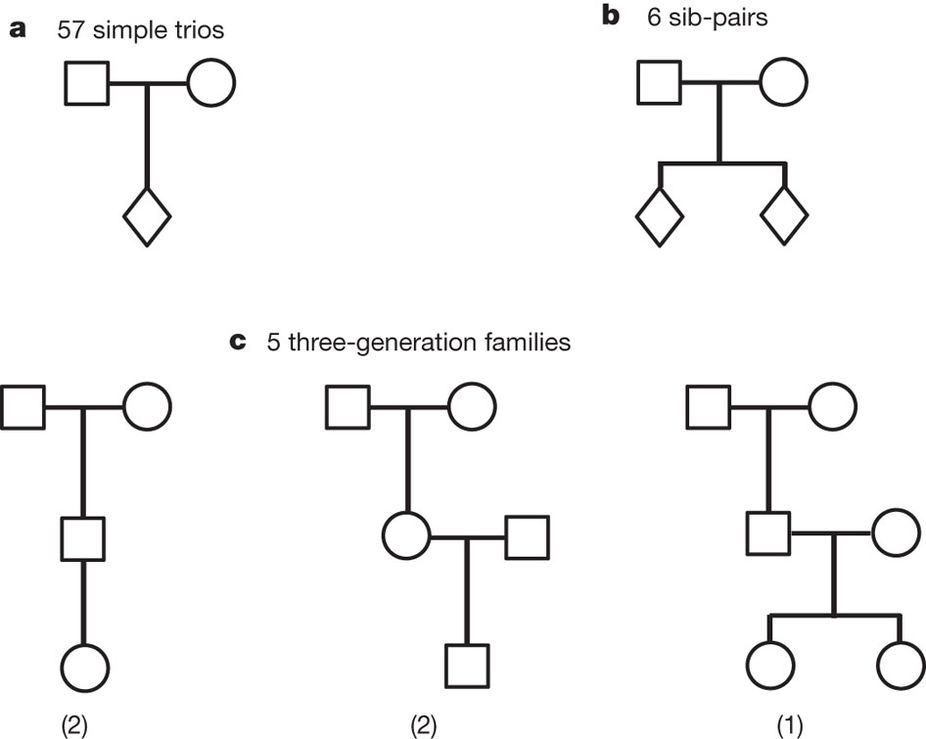
Kong et al, Nature, 2012
Table. De novo mutations observed with parental origin assigned (Kong et al, Nature, 2012).
| Trio | Father’s age | Mother’s age | Paternal chromosome | Maternal chromosome | Combined |
|---|---|---|---|---|---|
| Trio 1 | 21.8 | 19.3 | 39 | 9 | 48 |
| Trio 2 | 22.7 | 19.8 | 43 | 10 | 53 |
| Trio 3 | 25.0 | 22.1 | 51 | 11 | 62 |
| Trio 4 | 36.2 | 32.2 | 53 | 26 | 79 |
| Trio 5 | 40.0 | 39.1 | 91 | 15 | 106 |
| Mean | 29.1 | 26.5 | 55.4 | 14.2 | 69.6 |
| s.d. | 8.4 | 8.8 | 20.7 | 7.0 | 23.5 |
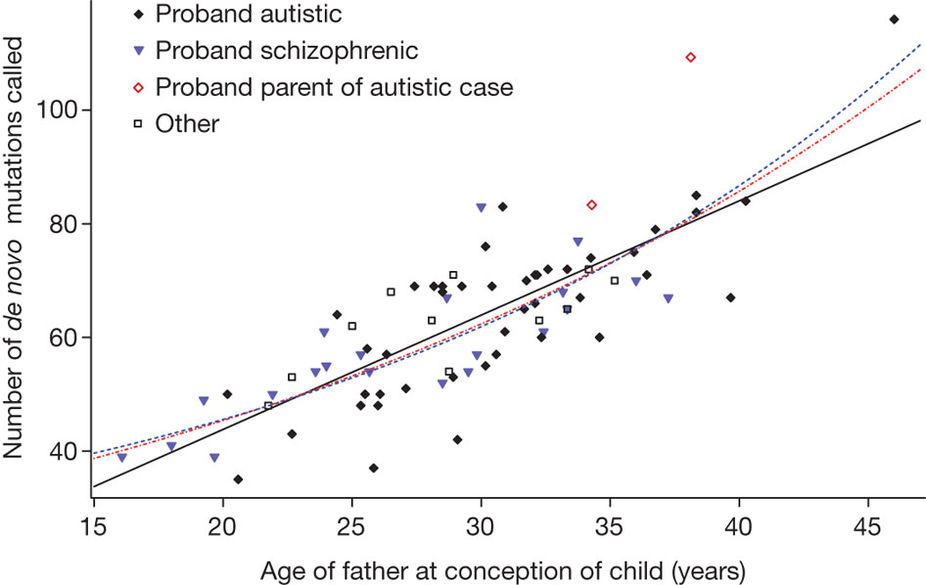
Kong et al, Nature, 2012
Fitting
Goal Of Fitting
- Fitting is the process of comparing a model to a compendium of data
- After fitting, we will have a model that explains existing data and can predict new data
Process of Fitting
The process of fitting is choosing parameter values so that a model explains the observed data as well as possible.
The key factor is how one defines the problem—i.e., how the distribution is described.
Three ingredients of a fitting problem
- Model form: What relationship do we assume between inputs and outputs?
- Loss / objective: What does it mean for the model to fit the data well?
- Assumptions: What statistical assumptions make the fit interpretable?
Caveats
- Any fitting result is highly dependent upon the correctness of the model
- Successful fitting requires concordance between the model and data
- Too little data and a model is underdetermined
- Unaccounted for variables can lead to systematic error
Since all models are wrong the scientist cannot obtain a “correct” one by excessive elaboration. On the contrary following William of Occam he should seek an economical description of natural phenomena. Just as the ability to devise simple but evocative models is the signature of the great scientist so overelaboration and overparameterization is often the mark of mediocrity. ~George Box, J American Stat Assoc, 1976
Any Fitting Depends on Model Correctness
Tyler Vigen. Spurious correlation.
Fitting does not happen in a vacuum!
Ordinary Least Squares
Ordinary Least Squares (OLS)
- Probably the most widely used estimation technique.
- Closely connected to maximum likelihood under a Gaussian noise model.
- Model assumes the output quantity is a linear combination of inputs.
Model setup
If we have a vector of \(n\) observations \(\mathbf{y}\), our predictions are going to follow the form:
\[ \mathbf{y} = \mathbf{X} \boldsymbol{\beta} + \boldsymbol{\epsilon} \]
Here:
- \(\mathbf{X}\) is a \(n \times p\) structure matrix (a.k.a. design matrix)
- \(\boldsymbol{\beta} \in \mathbb{R}^p\) is the vector of model parameters (regression coefficients)
- \(\boldsymbol{\epsilon} = \left( \epsilon_1, \epsilon_2, ... \epsilon_n \right)^\top\) is the noise present in the model
- Usual assumptions: uncorrelated and have constant variance \(\sigma^2\):
\[ \boldsymbol{\epsilon} \sim \left( \mathbf{0}, \sigma^2 \mathbf{I} \right) \]
Intuition
OLS picks the line, plane, or hyperplane that makes predictions as close as possible to the observed data in total squared distance.
Single variable case
For a single input variable, ordinary least squares is just:
\[ \mathbf{y} = m \mathbf{x} + \beta_0 + \boldsymbol{\epsilon} \]
Here, \(m\) is the slope, \(\beta_0\) is the intercept, and \(\boldsymbol{\epsilon}\) is the noise.
Multiple variable case
The structure matrix is little more than the data, sometimes transformed, usually with an intercept (offset). So, another way to write:
\[ \mathbf{y} = \mathbf{X} \boldsymbol{\beta} + \boldsymbol{\epsilon} \]
would be:
\[ \mathbf{y} = m_1 \mathbf{x}_1 + m_2 \mathbf{x}_2 + \ldots + \beta_0 + \boldsymbol{\epsilon} \]
The values of \(m\) and \(\beta_0\) that minimize the sum of squared error (SSE) are optimal.
Including the intercept term
Consider including the intercept \(\beta_0\) by augmenting the design matrix \(\mathbf{X}\) and parameter vector \(\boldsymbol{\beta}\):
\[ \mathbf{X} \boldsymbol{\beta} = \left(\mathbf{1} \; \mathbf{x}_1 \; \mathbf{x}_2 \; \cdots \; \mathbf{x}_p \right) \begin{pmatrix} \beta_0 \\ \beta_1 \\ \beta_2 \\ \vdots \\ \beta_p \end{pmatrix} \]
Now, the standard notation considers the intercept.
\[ \mathbf{y} = \mathbf{X} \boldsymbol{\beta} + \boldsymbol{\epsilon} \]
Least-squares objective and closed-form solution
Gauss and Markov in the early 1800s identified that the least squares estimate of \(\boldsymbol{\beta}\), \(\hat{\boldsymbol{\beta}}\), is obtained by minimizing the total squared error:
\[ \hat{\boldsymbol{\beta}} = \arg\min_{\boldsymbol{\beta}}{\left\Vert \mathbf{y} - \mathbf{X} \boldsymbol{\beta} \right\Vert^{2}} \]
When \(\mathbf{X}^\top \mathbf{X}\) is invertible, this can be directly calculated by:
\[ \hat{\boldsymbol{\beta}} = \mathbf{S}^{-1} \mathbf{X}^\top \mathbf{y} \]
where
\[ \mathbf{S} = \mathbf{X}^\top \mathbf{X} \]
Ordinary Least Squares and Maximum Likelihood
Why does least squares make sense?
\[ \log L(\boldsymbol{\beta},\sigma^{2}) = -\frac{n}{2} \log(2\pi\sigma^{2}) - \frac{1}{2 \sigma^{2}} \sum_{i=1}^{n} (y_{i}-\mathbf{x}_{i}^\top \boldsymbol{\beta})^{2} \]
Under the Gaussian noise assumption, maximizing the likelihood is therefore equivalent to minimizing:
\[ \sum_{i=1}^{n} (y_{i}-\mathbf{x}_{i}^\top \boldsymbol{\beta})^{2} \]
So least squares is also the maximum likelihood estimate under this model.
Statistical properties of the estimator
\(\hat{\boldsymbol{\beta}}\) is the maximum likelihood estimate of \(\boldsymbol{\beta}\), and has covariance \(\sigma^2 \mathbf{S}^{-1}\):
\[\hat{\boldsymbol{\beta}}\sim{}N\left(\boldsymbol{\beta},\sigma^2 \mathbf{S}^{-1}\right)\]
In the normal case (when our assumptions hold), \(\hat{\boldsymbol{\beta}}\) is an unbiased estimator of \(\boldsymbol{\beta}\). Making these calculations tractable for larger data sets used to be a challenge but is now trivial.
Advantages and limitations
When should we use ordinary least squares (OLS)?
- What are some of the assumptions?
- What are the implications of these assumptions not holding?
- What are some downsides?
Implementation
scikit-learn provides a very basic function for ordinary least squares.
scikit-learn
sklearn.linear_model.LinearRegressionfit_intercept: Should an intercept value be fit?- Centering or scaling the input variables should typically be done as a separate preprocessing step
- No tests for significance/model performance included.
- We’ll discuss evaluating the model in depth later.
NumPy
Or there’s an even more bare-bones function in NumPy: numpy.linalg.lstsq.
- Takes input variables
AandB. - Solves the equation \(Ax=B\) by computing a vector \(x\) that minimizes the Euclidean 2-norm \(\lVert B-Ax \rVert^2\).
Example
from sklearn.linear_model import LinearRegression
from sklearn.datasets import load_diabetes
from matplotlib import pyplot as plt
lr = LinearRegression()
data = load_diabetes()
y = data.target
lr.fit(data.data, y) # X, y
predicted = lr.predict(data.data)
fig, ax = plt.subplots()
ax.scatter(y, predicted, edgecolors=(0, 0, 0))
ax.plot([y.min(), y.max()], [y.min(), y.max()], 'k--', lw=4)
ax.set_xlabel('Measured')
ax.set_ylabel('Predicted')This plot shows training-set fit only. It is useful for illustration, but it does not tell us how well the model will generalize to new data.
Non-Linear Least Squares
Non-Linear Least Squares
Non-Linear Least Squares makes similar assumptions to ordinary least squares, but for arbitrary functions. Thus, instead of following the form:
\[ \mathbf{y} = \mathbf{X} \boldsymbol{\beta} + \boldsymbol{\epsilon} \]
Our input-output relationship is:
\[ \mathbf{y} = f(\mathbf{X}, \boldsymbol{\beta}) + \boldsymbol{\epsilon} \]
for the same construction of \(\boldsymbol{\epsilon} \sim \left( \mathbf{0}, \sigma^2 \mathbf{I} \right)\).
Transformation
- Non-linear least squares used to be mostly performed by transforming one’s data into a linear model.
- For instance, by taking the ratio of variables, or log-transforming them.
- This is now considered bad practice.
- Why?
Why move to non-linear least squares?
If we believe the underlying relationship is non-linear (e.g., curved), it is usually better to fit that curved model directly than to force the data into a linear form by transformation.
Non-Linear Least Squares
Algorithms
We again need to solve for \(\boldsymbol{\beta}\) to minimize the sum of squared error (SSE):
- There are many methods to solve these problems, and finding the true minimum is not a trivial task.
- We’re not going to cover how these algorithms work in depth.
Non-Linear Least Squares
Algorithms
One property we can take advantage of is that the gradient of the SSE with respect to \(\boldsymbol{\beta}\) at the minimum is zero (\(r_i\) is the residual of the \(i\)th point):
\[ {\frac {\partial S}{\partial \beta_j}}=2\sum_i r_i{\frac {\partial r_i}{\partial \beta_j}}=0 \]
- \(\frac{\partial r_i}{\partial \beta_j}\) is a function of both the non-linear function and the data.
- This can be expanded out through a first-order Taylor approximation.
- Doing so essentially performs ordinary least squares around the current point, for the linearized function.
\[ {\frac {\partial S}{\partial \beta_j}}=2\sum_i r_i{\frac {\partial r_i}{\partial \beta_j}}=0 \]
- \(\frac{\partial r_i}{\partial \beta_j}\) is a function of both the non-linear function and the data.
- This can be expanded out through a first-order Taylor approximation.
- Doing so essentially performs OLS around the current point.
- \({\frac{\partial r_i}{\partial \beta_j}}= -J_{ij}\), where \(J\) is the Jacobian of the function.
- Many non-linear least-squares solvers require \(J\) for this reason; it can be approximated by finite differences.
- Probably the most common method, Gauss-Newton, uses this property with Newton’s method.
Non-Linear Least Squares
Algorithms - Key Takeaways
- Unlike ordinary least squares, no guarantee about finding the optimal solution.
- Depending upon the data and model, there may be many local minima.
- This can still be interpreted in terms of shifting normal distributions around the data.
Implementation
SciPy’s scipy.optimize.least_squares is a very capable implementation.
- The main necessary parameters are:
fun, the functionx0, an initial guess for the parameter values
- Note that
funshould return a vector of the residuals- So it should handle all the data itself
NNLS Example
Binding Data
Let’s say we’re looking at a protein-protein interaction such as this:
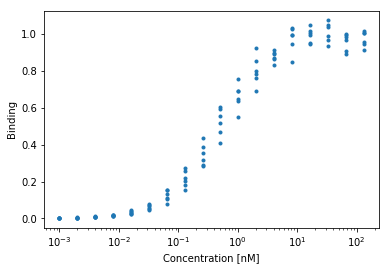
Let \(X\) denote ligand concentration and let \(\beta = k_1\) denote the association constant. We can predict the amount of binding we’d observe from a single-site binding model.
\[ y = f(X, \beta) + \epsilon \]
where,
\[ f(X, \beta) = \frac{X\beta}{1 + X\beta} \]
Implementation
We can define custom function: \(f([L], k) = \frac{k[L]}{1 + k[L]}\)
def klotz1(k1, lig):
"""
Calculates the fractional binding for a single-site receptor model.
Args:
k1: The association constant. Higher values indicate stronger binding affinity.
lig: The concentration of the free ligand.
Returns:
The predicted fractional occupancy (Y).
This will be a value strictly between 0.0 (no binding) and 1.0 (100% saturation).
"""
return (k1*lig)/(1 + k1*lig)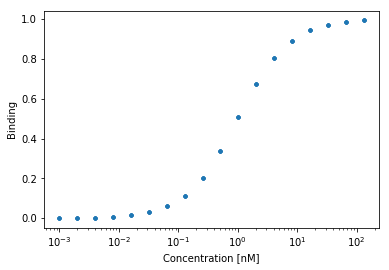
NNLS Example
Binding Data
SciPy asks for the residuals at each fitting point, so we need to convert a prediction to that:
sp.optimize.least_squares(ls_obj_k1, 1., args=(X,Y))
# --------
active_mask: array([ 0.])
cost: 0.0086776496708916573
fun: array([ 4.79e-05, 9.00e-05, -1.09e-04,
8.04e-04, -9.67e-04, 3.85e-03,
4.61e-03, 2.34e-03, 2.36e-02,
9.64e-03, -2.48e-02, 1.93e-02,
-4.93e-02, 5.54e-02, -3.66e-02,
2.97e-03, 3.39e-02, -8.74e-02])
grad: array([ -9.57228474e-09])
jac: array([[-0.00099809],
[-0.00199235],
[-0.0039695 ],
# ...
[-0.01608763],
[-0.00817133]])
message: '`gtol` termination condition is satisfied.'
nfev: 4
njev: 4
optimality: 9.5722847420895082e-09
status: 1
success: True
x: array([ 0.95864059])How to select the right model?
Generalized Linear Model
What if the error term isn’t Gaussian?
- In many cases linear regression can be inappropriate
- E.g. A measurement that is Poisson distributed
- E.g. A binary outcome
Logistic regression
- Commonly used to model binary outcomes.
- \(p(\mathbf{x}) = \frac{1}{1 + \exp\left(-\left(\beta_0 + \boldsymbol{\beta}^{\top}\mathbf{x}\right)\right)}\)
- Can we simply use logistic regression to analyze the Binding Data?
- Why or why not?
Can/should we use logistic regression for binding data?
- The choice depends on what you know about the system.
- If you have domain knowledge about receptor-ligand binding, a mechanistic model is usually preferable because its parameters are interpretable.
- We can use logistic regression for binding data.
- There is a connection: the Klotz binding model can be written in a logistic-like form.
The Klotz binding model and logistic regression
\[ \begin{aligned} y &= \frac{k[L]}{1 + k[L]} = \frac{1}{1 + (k[L])^{-1}} \\ &= \frac{1}{1 + \left( \exp(\ln(k)) \cdot \exp(\ln([L])) \right)^{-1}} \\ &= \frac{1}{1 + \exp \left( -\left( \ln(k) + \ln([L]) \right) \right)} \end{aligned} \]
- Logistic regression with the logarithm of ligand concentration, \(\ln([L])\), is related to the Klotz binding model.
- The scaling and transformation of the variables matters.
How to select the right model?
- If you have domain knowledge about the system, use it.
- Mechanistic models can give interpretable parameter estimates.
- If you do not have a mechanistic model, use a statistical model whose assumptions are reasonable.
- Off-the-shelf models often have good documentation, toy examples, and efficient implementations.
- Choose a model whose outputs answer your scientific question.
- Good prediction alone is not always enough if the goal is interpretation.
Review
Questions
- Given the binding data presented here, do you think a least squares model is most appropriate?
- How might you test whether your data seems to follow the assumptions of your model?
Reading & Resources
Review Questions
- Are OLS or NNLS guaranteed to find the optimal solution?
- How are new points predicted to be distributed in OLS?
- How are new points predicted to be distributed in NNLS?
- How might you determine whether the assumptions you made when running OLS are valid? (Hint: Think about the tests from lecture 1.)
- What is a situation in which the statistical assumptions of OLS can be valid but calculating a solution fails?
- You’re not sure a function you wrote to calculate the OLS estimator (Gauss-Markov) is working correctly. What is another relationship you could check to make sure you are getting the right answer?
- You’ve made a monitor that uses light scattering at three wavelengths to measure blood oxygenation. Design a model to convert from the light intensities to blood oxygenation using a set of calibration points. What is the absolute minimum number of calibration points you’d need? How would you expect new calibration points to be distributed?
- A team member suggests that the light-oxygenation relationship from (7) is log-linear instead of linear, and suggests using log(V) with OLS instead. What would you recommend?